Linked Life Data (LLD) is a data-as-a-service platform that provides access to 25 public biomedical databases through a single access point. The service allows writing of complex data analytical queries, answering complex bioinformatics questions such as 'give me all human genes located in Y-chromosome with the known molecular interactions'; simply navigate through the information, or export subsets like 'all approved drugs and their brand names'.
The service offers two different access levels:
| Feature | LLD Public | LLD Enterprise |
|---|---|---|
| SPARQL Endpoint | Limited to queries executed in 30s | Unlimited and exclusive server access |
| Secure HTTPS access | No | Yes |
| Write access | No | Yes |
| Data sources | copyright limited and copyright free sources | sources with commercial license |
| Updates | Annual | Monthly |
| Subscription fee | No | Yes |
| Professional services to improve the data | No | Yes |
| Support | None | Commercial support regulated by service level agreement |
We provide enterprise support of the linked data cloud. To contact us, please send an email to life-sciences@ontotext.com.
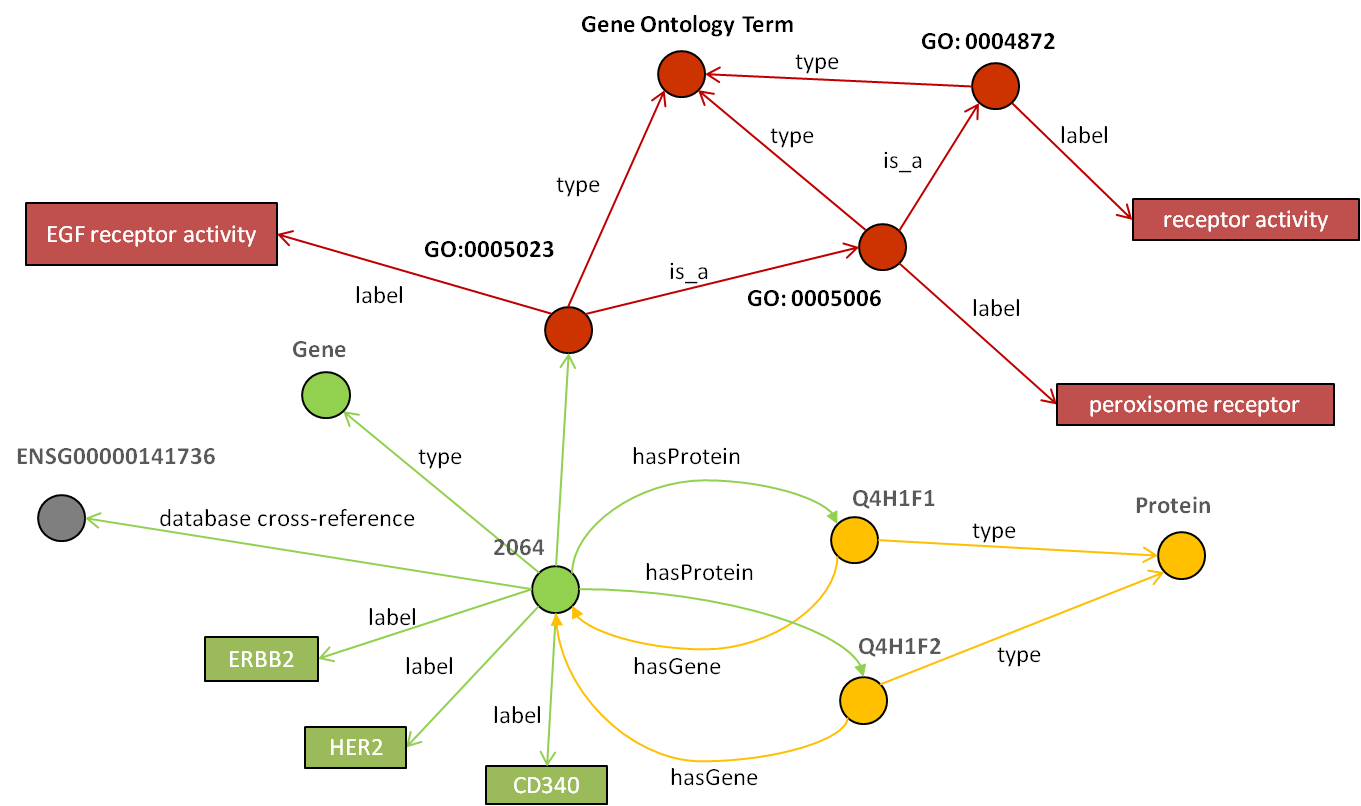
LLD uses a distributed graph data model to represent complex heterogeneous information. Imagine it as linked data. Linked data is method of publishing structured data so that it can be easily interlinked. The picture below demonstrates in different colours (i.e. different physical locations) how the information is represented and identified by global identifiers (URI). To create linked data you have to follow these few very simple steps:
GraphDB is a semantic repository - a software component for storing and manipulating huge quantities of RDF and linked data. This is the database instance used to power every LLD server node. More specifically, every node operates GraphDB-SE. GraphDB-SE is suitable for handling massive volumes of data and very intensive querying activities. It is designed as an enterprise-grade database management system. This has been made possible through:
GraphDB allows the loading of all 10 billion RDF statements in a single machine, and guarantees very fast query response time. The database also supports federated queries and links to URIs hosted by external systems.
In the process of integrating the 25 databases, which are part of the public service, the following linked data generation conventions were used:
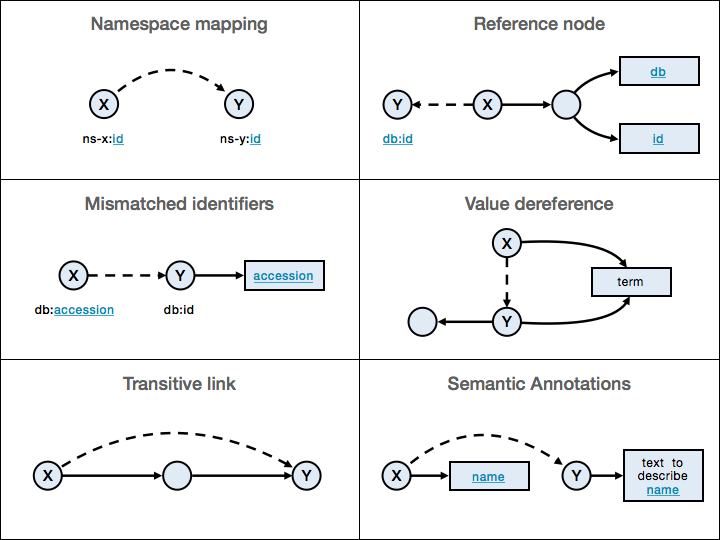
After all data is represented into RDF data format, many additional connections between the resources must be made, before it becomes truly “linked” data. The blue lines and the blue text of the captions (used either as part of the URI, or as literals) mark the criteria for linking the information. The specified mapping rules are applied only to the specified subsets of information. Another form of connection is Semantic Annotation, which uses NLP analysis to generate links between entities and unstructured textual fragments.
The service covers the full path of data - gene, protein, molecular interaction, pathway, target, drug, disease and clinical trial related information. These are the primary entities of a knowledge base composed by structured databases (NCBI Gene, Uniprot, DrugBank, BioPAX and many more), terminologies (UMLS, OBO), and semi-structured documents (Pubmed, ClinicalTrials.gov). The public service integrates only free databases. While others, used in the Enterprice edition have further license restrictions. To check the complete list, visit the current list of processed databases.
This work was partially funded by Linked Life Data and the EU IST projects, KHRESMOI (FP7-257528) and LarKC (FP7-215535)